A University of Queensland research group has already changed the way it works due to a new service added to high-performance computer Bunya.
Associate Professor Rodrigo Suarez and Postdoctoral Research Fellow Dr Dylan Anthony Black of UQ’s School of Biomedical Sciences say onBunya, a version of Open OnDemand customised to the HPC, has altered the way their research group, which studies brain evolution and development, integrates data visualisation and compute.
The Suárez laboratory extensively uses Bunya and onBunya to aid in the analysis and visualisation of its large neurobiology datasets.
OnBunya, involving remote web access and an easier graphical user interface (GUI) for Bunya, became operational earlier this year for all of the HPC’s users after more than six months of user testing.
OnBunya provides researchers with an easily accessible interface to the powerful hardware Bunya has to offer. The first and current iteration of onBunya also makes it easy to run GUI software on the HPC.
“The addition of onBunya confers several benefits,” said Dr Black. “For our experienced users, onBunya allows for critical visual inspection of data, and integration of GUI-dependent steps into our workflows.
“Importantly, onBunya is also allowing our lab members with less experience to access high-performance computing systems, lowering the barrier to entry, and democratising access to these resources.”
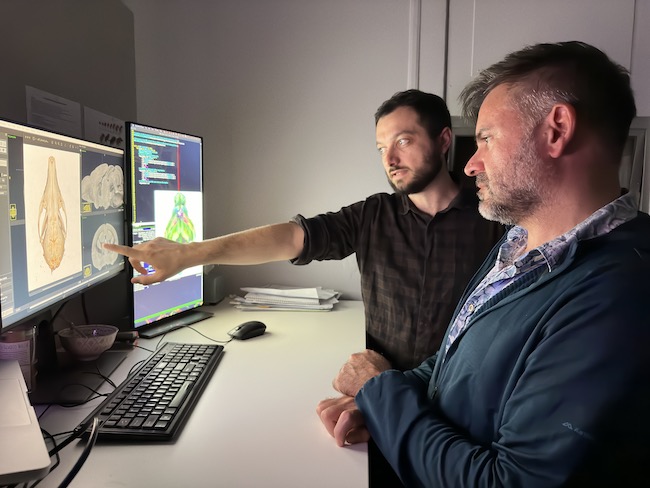

Broadly, the Suárez group’s work is focused on the evolution, development, and maturation of mammalian brain structures, such as the cerebral cortex. A specific focus is on the initiation and development of electrical activity in cells of the cortex, a brain structure unique to mammals, including humans.
The researchers believe that better understanding the evolutionary origins of mammalian nervous systems, and how these systems are built via complex developmental programs and interactions with the environment, will provide critical insight into healthy brain formation.
“In our lab, we are interested in knowing where electrical activity begins in the developing cortex, how it relates to the emerging brain anatomy and sensory systems, and what important functions it serves that leads to a healthy mature brain,” said Dr Black.
“One part of this work involves observing signals occurring in the living brain. Recording this activity over large brain areas, and long time periods, produces enormous data files. Simultaneously, we take recordings that help us remove the influence of artifacts, such as blood flow and animal movements, which alters neural activity readouts, and improve the signal-to-noise ratio in our data.”
The group uses onBunya to facilitate an efficient workflow by seamlessly integrating the steps for distinguishing neural signals from artifacts. “Our experience indicates that other ‘human-in-the-loop’ [computer models requiring human interaction] workflows stand to gain from similar integration, enhancing efficiency,” said Dr Black.
“Processing this data all together and preparing it for analysis requires not only rapid access to these files but also the memory and compute to perform steps such as image correction, normalisation, and dimensionality reduction.”
Another goal of the group’s work is to better understand the relationship between the brain’s electrical activity and the underlying anatomical connectivity. “We use varied techniques to do this, including diffusion-weighted magnetic resonance imaging that allows us to examine white matter tracts – the insulated fibres that connect different brain areas.” (See Figure 1 below.)
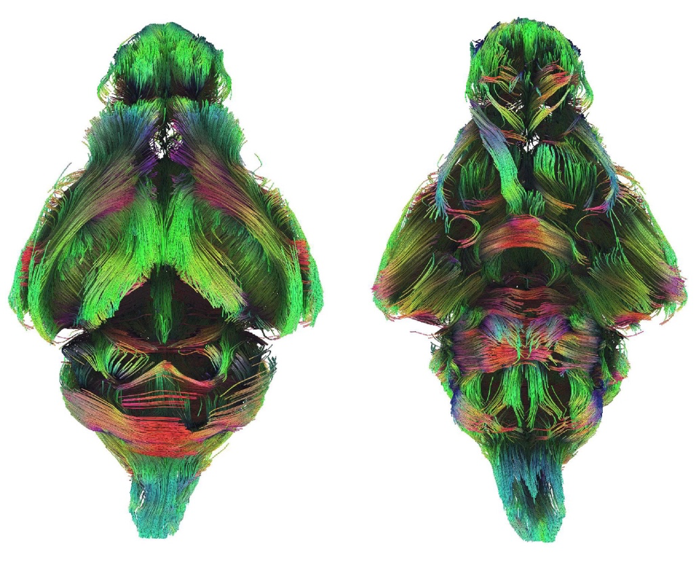
“Our questions range from the developmental mechanisms responsible for the evolutionary origin of complex trait innovations, to information processing across brain regions,” said Associate Professor Suárez.
“Advancing our understanding of how mammalian brains develop has historically been complicated by the fact that much brain development occurs in utero, which in turn makes it difficult to study.”
The Suárez group has used brain scans of Australian marsupial the fat-tailed dunnart in its work.
“Marsupials help overcome these complications as they give birth to extremely underdeveloped young. This provides unprecedented access to key developmental events early in life, in a way similar to what is possible in zebrafish or fruit flies but in a mammalian species with a cerebral cortex similar to humans.
“We are confident that combining this model of mammalian brain development with state-of-the-art methods and analytical techniques will yield insights into general principles governing how remarkable biological structures such as brains form, mature, and diversify, as well as how developmental mechanisms can go astray in neurodevelopmental disorders, such as in autism spectrum disorder,” said Associate Professor Suárez.
The Suárez group use supercomputer Bunya across nearly all of its research methods, including single-cell RNA sequencing, high-throughput histology imaging and quantification, in vivo microscopy of brain neurons firing in real time, machine learning pipelines, and magnetic resonance structural and diffusion-weighted imaging.
“A branch of our work is centred on optical and magnetic resonance imaging methods, which are inherently visual, and the ability to visualise our data via GUI interfaces [via onBunya] in a manner continuous with our processing pipelines has already been of great value to us,” said Dr Black.
“Moreover, visualisation of large data files is facilitated by graphical acceleration using high-end GPUs [graphics processing units], such as NVIDIA’s H100s.
“This data undergoes complex and demanding processing and analytical steps that demand high-end computational resources in order to be possible or practical,” said Dr Black.
Some aspects of the group’s research simply would not be possible without the Bunya infrastructure. “Our data regularly exceeds the memory limits of even high-end analytical workstations, and our compute requirements would mean that analysis on local workstations would take excessive amounts of time, severely impacting productivity and rate of progress.
“Much of our current research would be impossible without resources such as the Bunya cluster, and we would be required to look for compute externally at a significant cost.
“If we had to look externally for equivalent compute, this would also likely be without the thorough support that we receive internally via the staff at the Research Computing Centre. Productivity and feasibility of our work would be markedly impacted by the absence of this expertise.
“We interact daily with onBunya and regularly communicate with the RCC support team to enhance our capabilities by integrating new analytical and visualisation tools.”
One example Dr Black gave of how their research would be impacted without Bunya relates to the live brain imaging they perform. “To understand the range of brain activity patterns seen at early developmental stages, we must record this activity for significant periods of time. However, this quickly generates large datasets that exceed the memory limitations of common high-end workstations.
“Without access to Bunya, continuous recordings would not be possible and would have to be divided into smaller sessions with timing gaps that would impact interpretation of the data we collect.
“As more lab members engage with the Linux environment through onBunya’s user-friendly interface, our reliance on Bunya is expected to grow.
Other eResearch infrastructure
The Suárez group also depends on QRIScloud, the Queensland node of the ARDC Nectar Research Cloud that is jointly managed by RCC and QCIF.
In the background, QRIScloud uses UQ Research Data Manager’s (RDM) “Q” collections and UQ/RCC’s MeDiCI data fabric for the safe transport and long-term durable storage of large volumes of raw data that the group generates in the laboratory.
“Our storage needs exceed what would ever be feasible via local storage and would also be dramatically less secure and effectively backed up than what is provided by this infrastructure,” said Dr Black.
MeDiCI allows researchers to store their data, and almost immediately have it available for processing on the Bunya supercomputer, providing a powerful workflow to enable science at extreme scale.
More information about onBunya
For further information about onBunya, please read RCC’s article about Open OnDemand going live on Bunya, published on 14 March 2024.



